Introduction
NEREIDS (Neutron rEsonance REsolved Imaging Data analysis System) is a Rust-based library for neutron resonance imaging at the VENUS beamline, Spallation Neutron Source (SNS), Oak Ridge National Laboratory (ORNL).
What It Does
NEREIDS provides end-to-end analysis for time-of-flight (TOF) neutron resonance imaging: input hyperspectral TOF data, output spatially resolved isotopic and elemental composition maps.
The analysis pipeline:
- Load raw TOF imaging data (TIFF stacks, NeXus/HDF5, or pre-normalized transmission)
- Normalize sample and open-beam measurements to transmission
- Configure isotopes of interest using ENDF nuclear data
- Fit resonance models to measured transmission spectra
- Map fitted parameters (areal density, temperature) across each pixel
Three Deliverables
NEREIDS ships in three forms:
| Deliverable | Use case |
|---|---|
Rust library (nereids-* crates) | Embed in Rust applications, maximum performance |
Python bindings (pip install nereids) | Jupyter notebooks, scripting, integration with NumPy/SciPy |
Desktop GUI (nereids-gui) | Interactive analysis with visual feedback |
Relationship to SAMMY
NEREIDS implements the same physics as SAMMY (a Fortran code for multilevel R-matrix analysis of neutron data, ORNL/TM-9179/R8), rewritten in Rust with modern tooling. All physics modules reference specific SAMMY source files and equation numbers in their documentation.
Key formalisms from SAMMY:
- Reich-Moore R-matrix (LRF=3)
- Single-Level Breit-Wigner (LRF=1/2)
- R-Matrix Limited (LRF=7)
- Free Gas Model Doppler broadening
- Gaussian + exponential resolution broadening
- Unresolved Resonance Region (LRU=2)
Next Steps
- Install NEREIDS for your platform
- Try the Rust quickstart or Python quickstart
- Explore the GUI walkthrough for interactive analysis
Installation
Rust Library
Add the top-level orchestration crate (re-exports all lower-level crates):
[dependencies]
nereids-pipeline = "0.1"
Or add individual crates (nereids-core, nereids-endf, nereids-physics,
nereids-fitting, nereids-io) for finer-grained dependency control.
Requirements: Rust edition 2024 (rustc 1.85+).
Optional: HDF5 support
The nereids-io crate has an optional hdf5 feature for NeXus file support:
[dependencies]
nereids-io = { version = "0.1", features = ["hdf5"] }
This requires the HDF5 C library to be installed on your system.
Python Bindings
pip install nereids # available after first public release
Requirements: Python 3.10+ and NumPy.
Desktop GUI
macOS (Homebrew)
brew install --cask ornlneutronimaging/nereids/nereids # available after first public release
From Source
git clone https://github.com/ornlneutronimaging/NEREIDS.git
cd NEREIDS
cargo run --release -p nereids-gui
Building from source requires CMake (for HDF5) and a Rust toolchain.
Development Setup
For contributors working on NEREIDS itself:
git clone https://github.com/ornlneutronimaging/NEREIDS.git
cd NEREIDS
# Build everything
cargo build --workspace
# Run tests
cargo test --workspace --exclude nereids-python
# Build Python bindings (requires pixi)
pixi run build
pixi run test-python
See Contributing for the full development workflow.
Quickstart: Rust
This example loads ENDF resonance data for U-238, computes a theoretical transmission spectrum, and fits it to recover the areal density.
Setup
# Cargo.toml
[dependencies]
nereids-core = "0.1"
nereids-endf = "0.1"
nereids-physics = "0.1"
nereids-fitting = "0.1"
Load ENDF Data
use nereids_core::types::Isotope; use nereids_endf::retrieval::{EndfLibrary, EndfRetriever, mat_number}; use nereids_endf::parser::parse_endf_file2; use nereids_physics::transmission::{forward_model, SampleParams}; use nereids_fitting::lm::{levenberg_marquardt, LmConfig}; use nereids_fitting::transmission_model::TransmissionFitModel; use nereids_fitting::parameters::{FitParameter, ParameterSet}; fn main() -> Result<(), Box<dyn std::error::Error>> { // Download and cache ENDF/B-VIII.1 data for U-238 (Z=92, A=238) let isotope = Isotope::new(92, 238)?; let retriever = EndfRetriever::new(); let mat = mat_number(&isotope).expect("U-238 has a known MAT number"); let (_path, endf_text) = retriever.get_endf_file(&isotope, EndfLibrary::EndfB8_1, mat)?; let resonance_data = parse_endf_file2(&endf_text)?; println!("U-238: {} resonances, AWR = {:.1}", resonance_data.total_resonance_count(), resonance_data.awr);
Compute a Forward Model
#![allow(unused)] fn main() { // Energy grid: 1 to 30 eV (covers the 6.67 eV and 20.9 eV resonances) let energies: Vec<f64> = (0..2000) .map(|i| 1.0 + (i as f64) * 29.0 / 2000.0) .collect(); // Sample: U-238 at 0.001 atoms/barn, room temperature let sample = SampleParams::new(300.0, vec![(resonance_data.clone(), 0.001)])?; // No instrument resolution broadening for this example let transmission = forward_model(&energies, &sample, None)?; // transmission[i] is T(E_i) in [0, 1], with dips at resonance energies }
Fit a Measured Spectrum
#![allow(unused)] fn main() { // Simulate measured data (in practice, load from TIFF/NeXus) let measured_t = transmission.clone(); let sigma: Vec<f64> = vec![0.01; measured_t.len()]; // Set up the fit model: one density parameter at index 0 let model = TransmissionFitModel::new( energies.clone(), vec![resonance_data], 300.0, // temperature_k None, // no instrument resolution vec![0], // density_indices None, // no temperature fitting None, // no precomputed cross-sections )?; // Initial guess: density = 0.0005 atoms/barn (non-negative constraint) let mut params = ParameterSet::new(vec![ FitParameter::non_negative("U-238 density", 0.0005), ]); let config = LmConfig::default(); let result = levenberg_marquardt(&model, &measured_t, &sigma, &mut params, &config)?; println!("Fitted density: {:.6} atoms/barn", result.params[0]); println!("Reduced chi-squared: {:.3}", result.reduced_chi_squared); println!("Converged: {}", result.converged); Ok(()) } }
Next Steps
- See the API Reference for the full API
- Explore the Python quickstart for a NumPy-based workflow
Quickstart: Python
This example uses the NEREIDS Python bindings to load ENDF data, compute a forward model, and fit a transmission spectrum.
Setup
pip install nereids numpy matplotlib
Load ENDF Data and Compute Transmission
import nereids
import numpy as np
# Load ENDF/B-VIII.1 resonance data for U-238
u238 = nereids.load_endf(92, 238)
print(f"U-238: {u238.n_resonances} resonances, AWR = {u238.awr:.1f}")
# Energy grid: 1 to 30 eV
energies = np.linspace(1.0, 30.0, 2000)
# Compute transmission for 0.001 atoms/barn at 300 K
transmission = nereids.forward_model(
energies,
[(u238, 0.001)],
temperature_k=300.0,
flight_path_m=25.0,
delta_t_us=5.0,
delta_l_m=0.005,
)
Plot the Spectrum
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 4))
plt.plot(energies, transmission, linewidth=0.8)
plt.xlabel("Energy (eV)")
plt.ylabel("Transmission")
plt.title("U-238 Forward Model (0.001 at/barn, 300 K)")
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
Spatial Mapping
For imaging data (3D transmission arrays), use the typed API:
# transmission_3d: shape (n_energies, height, width)
# uncertainty_3d: shape (n_energies, height, width)
data = nereids.from_transmission(transmission_3d, uncertainty_3d)
result = nereids.spatial_map_typed(
data,
energies,
[u238],
temperature_k=300.0,
flight_path_m=25.0,
delta_t_us=5.0,
delta_l_m=0.005,
)
# result.density_maps[0] is a 2D array of U-238 areal density at each pixel
# result.converged_map shows which pixels converged
print(f"Converged: {result.n_converged}/{result.n_total} pixels")
For raw count data (Poisson-optimal fitting):
data = nereids.from_counts(sample_counts_3d, open_beam_counts_3d)
result = nereids.spatial_map_typed(data, energies, [u238])
Detectability Analysis
Check whether a trace isotope is detectable in a given matrix:
fe56 = nereids.load_endf(26, 56) # matrix: Fe-56
ag107 = nereids.load_endf(47, 107) # trace: Ag-107
report = nereids.trace_detectability(
matrix=fe56,
matrix_density=0.01,
trace=ag107,
trace_ppm=100.0,
energies=energies,
i0=1e6,
)
print(f"Detectable: {report.detectable}")
print(f"Peak SNR: {report.peak_snr:.1f} at {report.peak_energy_ev:.2f} eV")
Next Steps
- See the GUI walkthrough for interactive analysis
- Explore the Architecture chapter for the crate structure
- Browse the API Reference for the full Rust docs
GUI Walkthrough
The NEREIDS desktop application provides interactive neutron resonance imaging analysis with visual feedback at every step.
Launch
# Homebrew (macOS)
brew install --cask ornlneutronimaging/nereids/nereids
# Or pip
pip install nereids-gui
nereids-gui
# Or from source
cargo run --release -p nereids-gui
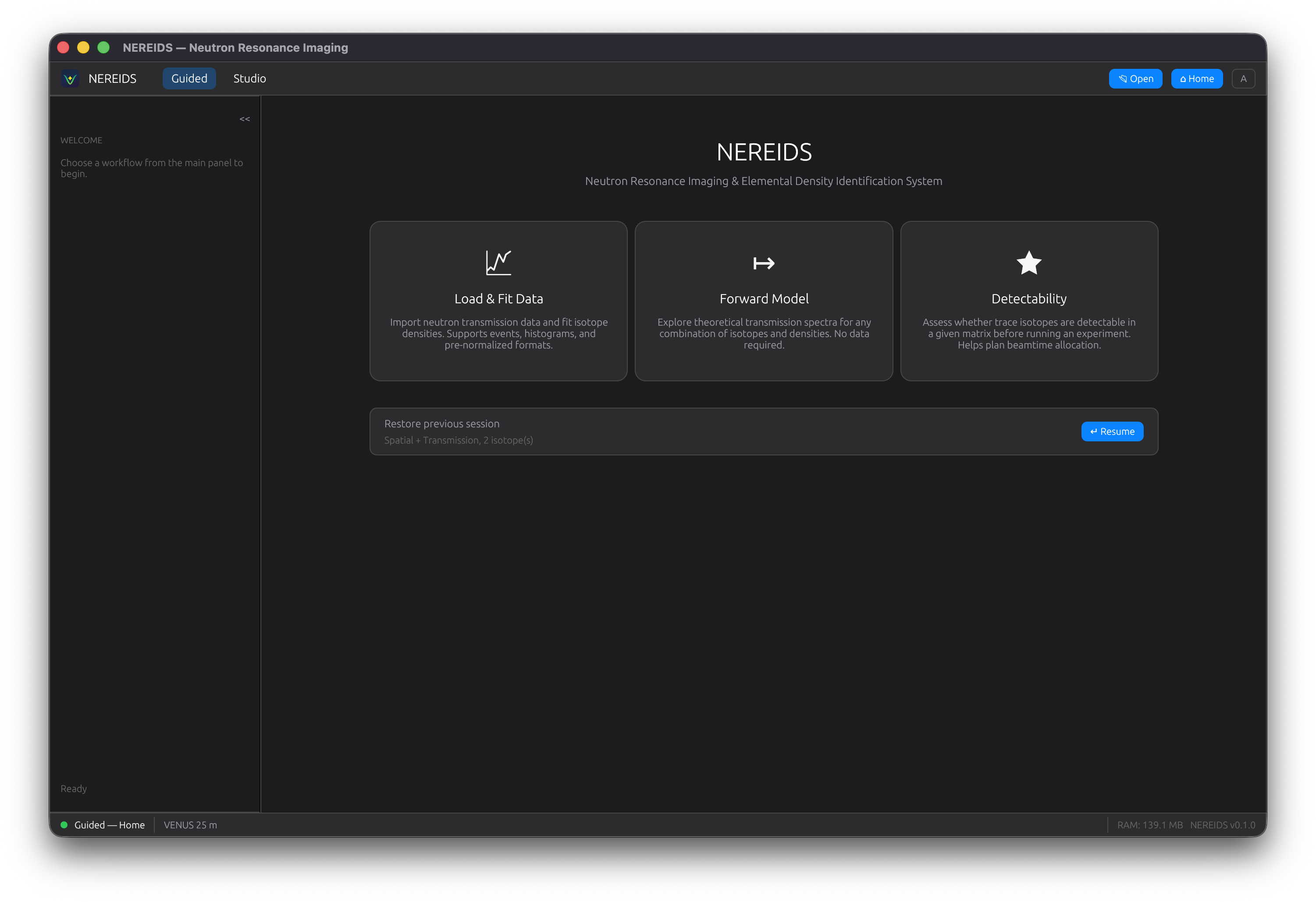
Landing Page
The landing page presents three entry points:
- Single Spectrum -- fit a single transmission spectrum to recover isotope densities
- Spatial Map -- fit every pixel in a transmission image stack
- Tools -- forward model, detectability analysis, periodic table

Decision Wizard
After selecting an entry point, a short wizard asks:
- Fitting type: Single spectrum or spatial map
- Data format: Raw events (NeXus), pre-normalized TIFF, or transmission TIFF
The wizard configures a dynamic pipeline with only the steps relevant to your data format. Six distinct pipelines are available.
Pipeline Steps
Load
Select sample data, open beam, and spectrum files. Supports multi-frame TIFF stacks, TIFF folders, and NeXus/HDF5 event data. The GUI auto-detects the file format and loads data when all fields are filled.
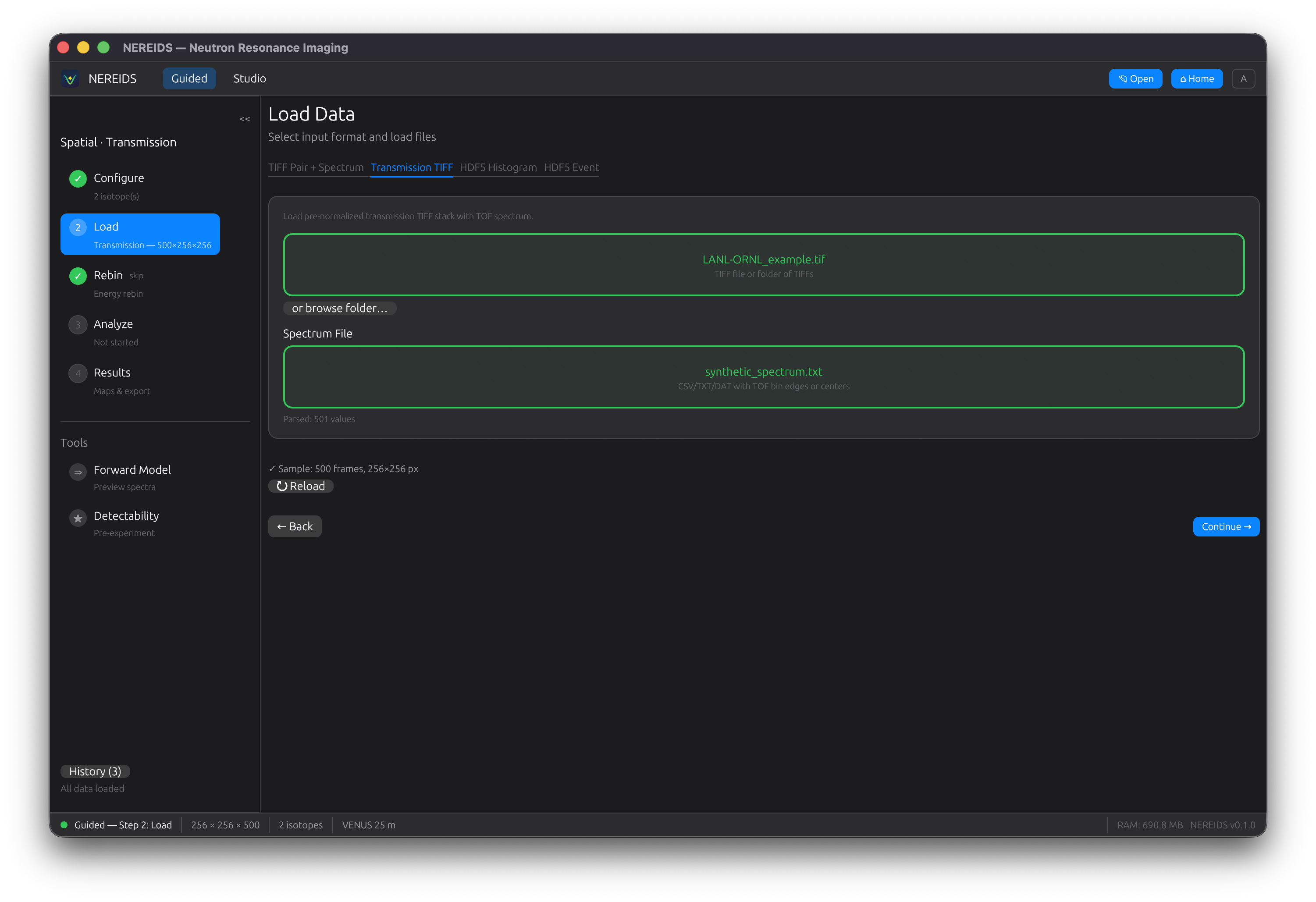
Normalize
For raw data pipelines (TIFF pair or NeXus events), the Normalize step computes transmission from sample and open-beam measurements. Pre-normalized and transmission TIFF pipelines skip this step automatically.
Configure
Select isotopes of interest from the periodic table. ENDF nuclear data is fetched automatically from IAEA servers and cached locally. Each isotope shows a status badge (Pending, Fetching, Loaded, Failed).
Configure beamline parameters (flight path, timing resolution) and solver settings (Levenberg-Marquardt or Poisson KL divergence).
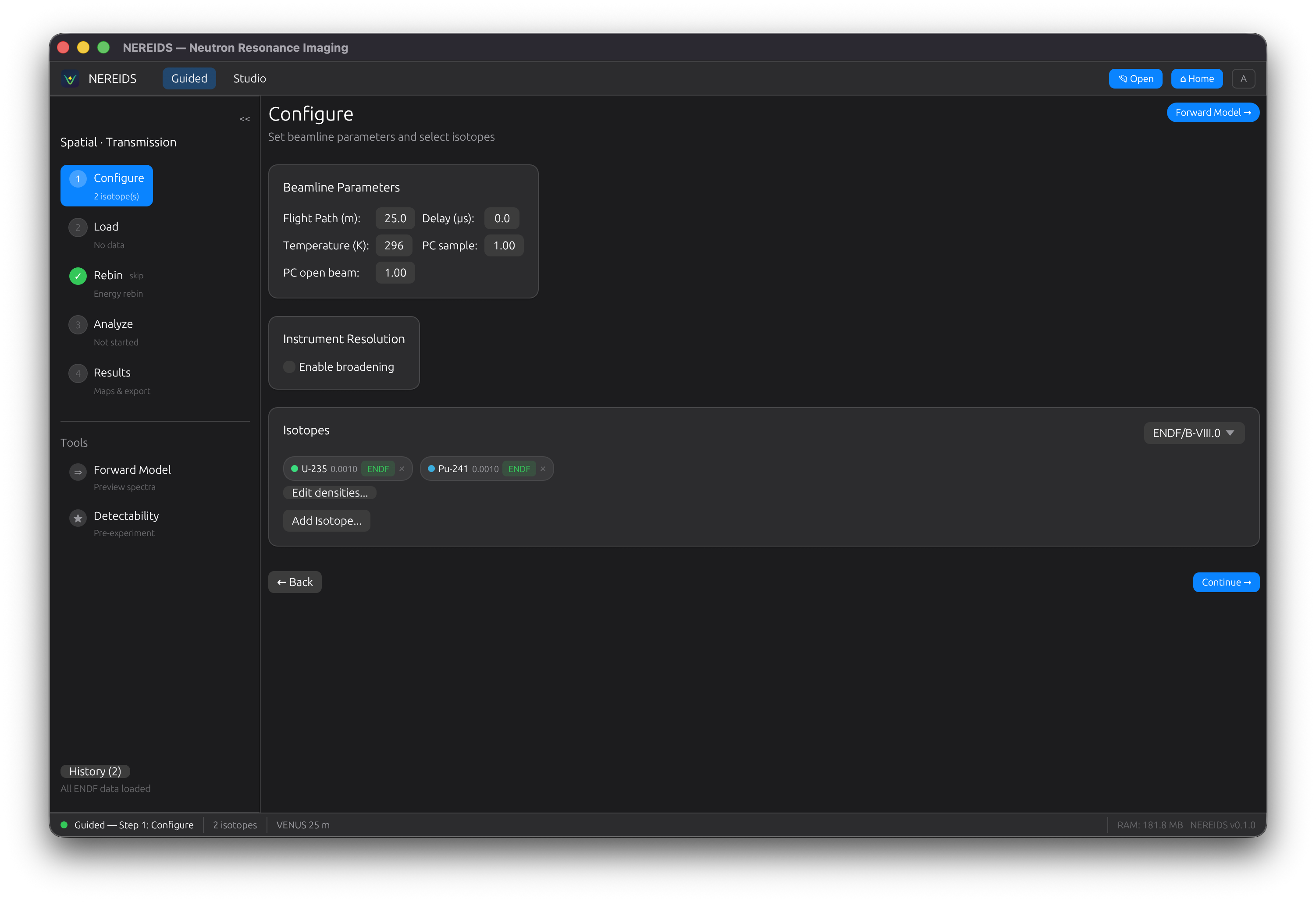
Analyze
Run the fit. For spatial maps, a progress bar tracks per-pixel fitting with rayon parallelism. Click any pixel to inspect its individual fit. Fit feedback shows green (good fit) or red (failed) status.
Draw regions of interest (ROI) with Shift+drag. Multiple ROIs are supported with move, select, and delete operations.
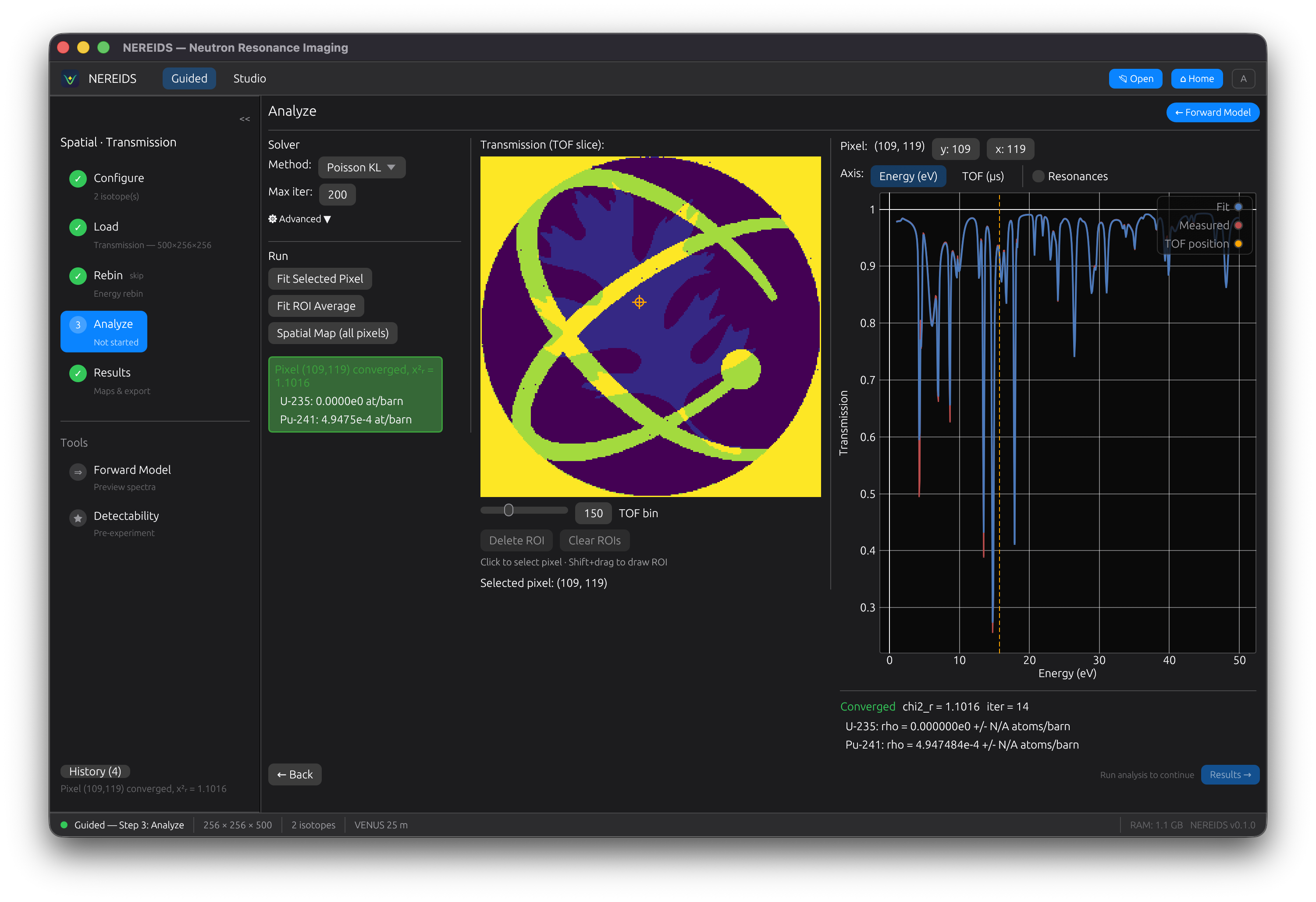
Results
View density maps for each fitted isotope. Summary statistics show convergence rate, median chi-squared, and isotope count. Open results in Studio for detailed exploration.
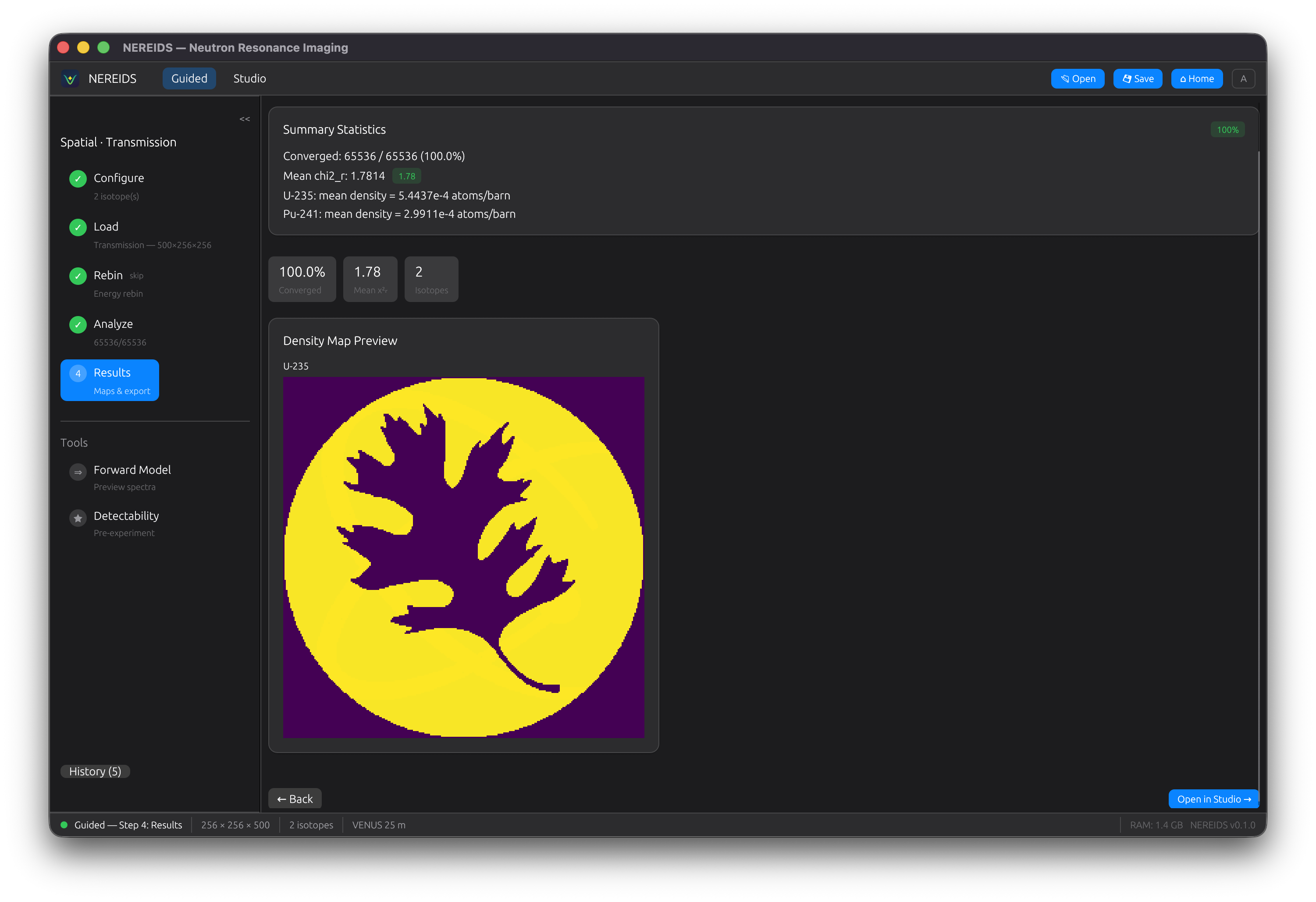
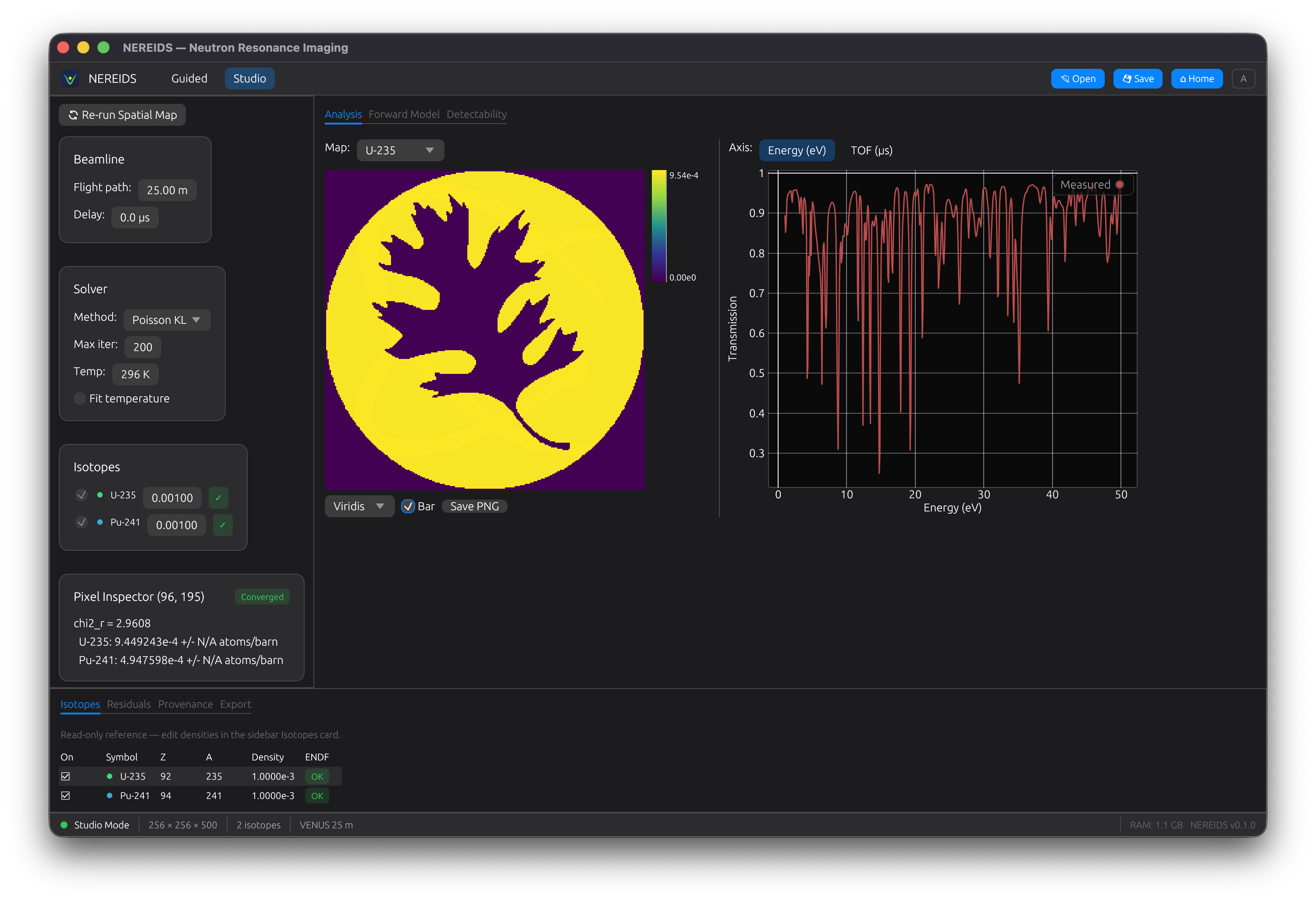
Studio Mode
Studio provides a "Final Cut"-style workspace for exploring results:
- Document tabs: switch between Analysis, Forward Model, and Detectability
- Main viewer: density map with colormap selection and colorbar
- Spectrum panel: click any pixel to see its fitted spectrum
- Bottom dock: Isotopes, Residuals, Provenance, and Export panels
- Inspector sidebar: per-pixel parameter values
Tools
Forward Model
Compute theoretical transmission spectra for arbitrary isotope mixtures. Adjust densities with sliders and see the spectrum update in real-time. Hero spectrum layout with per-isotope contribution lines.
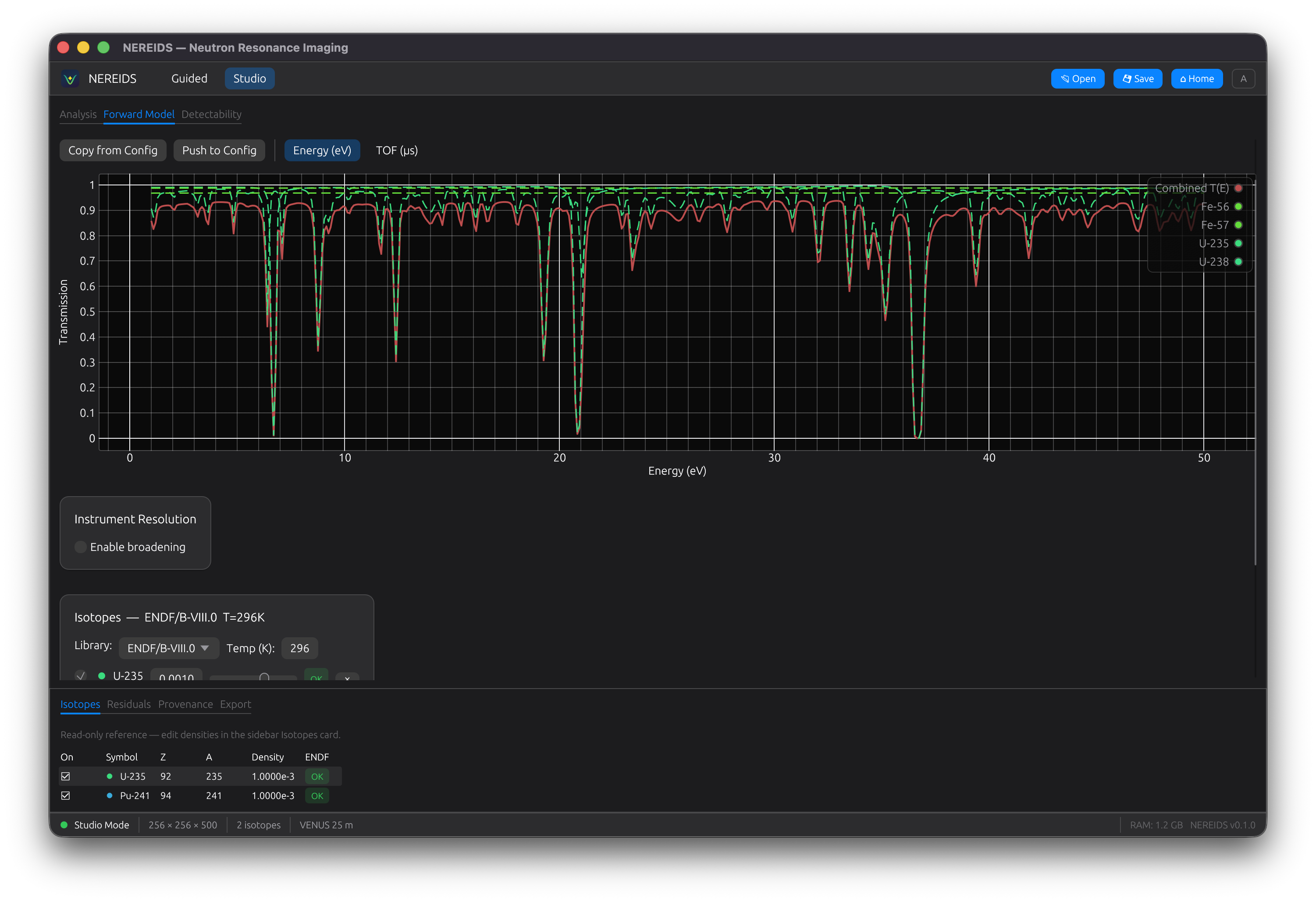
Detectability
Analyze whether a trace isotope is detectable in a given matrix material. Multi-matrix support with resolution broadening. Shows a delta-T spectrum and verdict badges (DETECTABLE / NOT DETECTABLE / OPAQUE MATRIX).
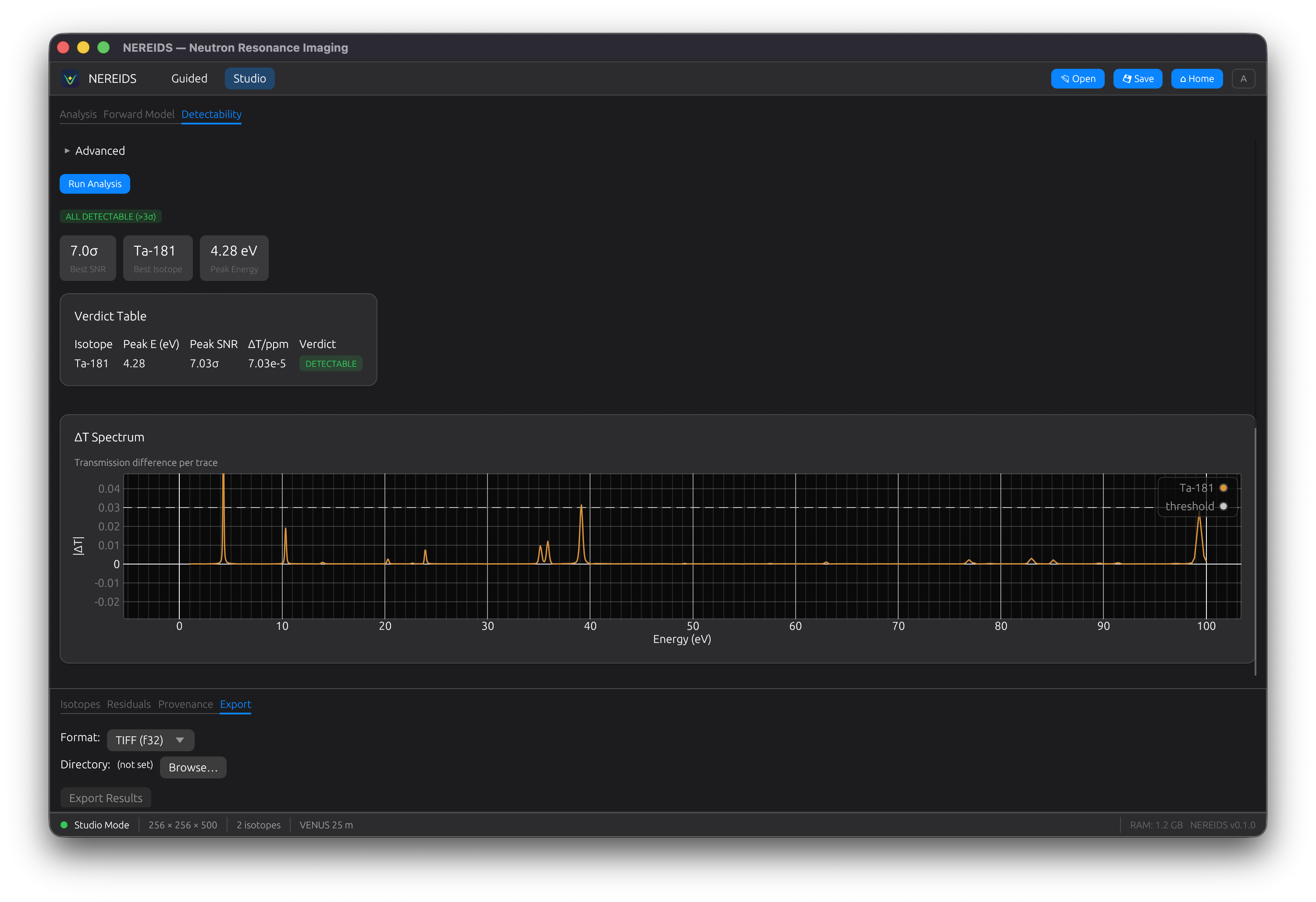
Periodic Table
Interactive 18-column periodic table for selecting isotopes. Click an element to see its natural isotopes with abundance percentages. Supports multi-select with density input. ENDF/B-VIII.0 availability hints shown for each isotope.
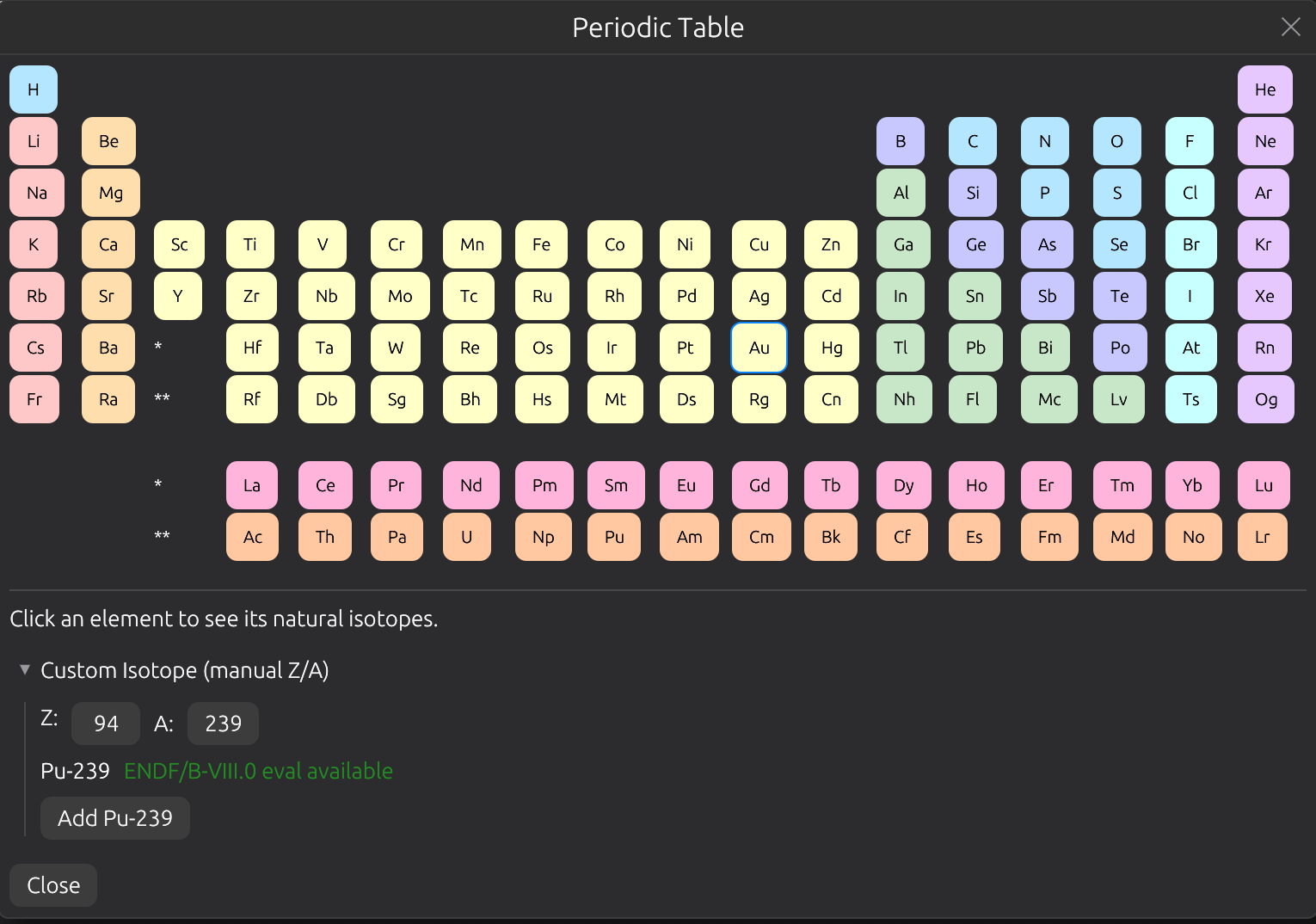
Project Files
Save and load analysis sessions as HDF5 project files (.nereids):
- Cmd+S (macOS) / Ctrl+S (Linux): quick-save
- File > Save: save with dialog
- File > Open: load a saved project
Project files store raw data, pipeline configuration, and results. Embedded data mode bundles everything into a single portable file.
Architecture
NEREIDS is organized as a Rust workspace with seven library crates, a GUI application, and Python bindings.
Crate Dependency Graph
endf-mat (standalone lookup tables)
|
nereids-core (types, constants)
/ | \
nereids-endf | nereids-io
(ENDF) | (TIFF, NeXus)
\ |
nereids-physics
(cross-sections)
\
nereids-fitting
(LM, Poisson)
|
nereids-pipeline ── nereids-io
(orchestration)
/ \
nereids-python nereids-gui
(PyO3 bindings) (egui desktop)
Crate Overview
| Crate | Purpose |
|---|---|
endf-mat | Zero-dependency lookup tables: element symbols, MAT numbers, natural abundances, ZA encoding |
nereids-core | Core types (Isotope, Resonance), physical constants, element data, error types |
nereids-endf | ENDF file retrieval from IAEA, local caching, File 2 resonance parameter parsing |
nereids-physics | Cross-section calculation (Reich-Moore, SLBW, RML, URR), Doppler/resolution broadening, Beer-Lambert transmission |
nereids-io | TIFF stack and NeXus/HDF5 loading, TOF-to-energy conversion, normalization, export |
nereids-fitting | Levenberg-Marquardt and Poisson KL divergence optimizers, transmission fit model |
nereids-pipeline | Single-spectrum fitting, per-pixel spatial mapping (rayon), trace detectability |
nereids-python | PyO3 Python bindings (not published to crates.io) |
nereids-gui | egui desktop application (not published to crates.io) |
Data Flow
The standard analysis pipeline processes data through these stages:
Raw TOF data (TIFF/NeXus)
│
▼
Normalization (sample / open_beam → transmission) [nereids-io]
│
▼
Energy conversion (TOF bin edges → energy centers) [nereids-io]
│
▼
ENDF data (fetch resonance parameters from IAEA) [nereids-endf]
│
▼
Forward model (cross-sections → broadening → T(E)) [nereids-physics]
│
▼
Fitting (minimize |T_measured - T_model|) [nereids-fitting]
│
▼
Spatial mapping (fit each pixel in parallel) [nereids-pipeline]
│
▼
Density maps, chi² maps, convergence maps [output]
Key Design Decisions
Exact SAMMY Physics
All physics modules implement the exact formalisms from the SAMMY Fortran code, with no ad-hoc approximations. Every module references specific SAMMY source files and equation numbers. See the Physics Reference for details.
Workspace Architecture
The workspace is structured so that each crate has a single responsibility
and minimal dependencies. nereids-core is the foundation with zero internal
dependencies. Higher-level crates compose lower-level ones.
See ADR 0001 for the full rationale.
Parallel Spatial Mapping
Per-pixel fitting uses rayon for data parallelism. The outer pixel loop runs on a dedicated thread pool to avoid deadlocking with inner parallel operations (cross-section calculation, broadening).
Python Bindings via PyO3
The Python bindings expose a high-level API (load_endf, forward_model,
fit_spectrum, spatial_map) that maps directly to the Rust pipeline.
NumPy arrays are zero-copy where possible via numpy crate integration.
Physics Reference
NEREIDS implements exact SAMMY physics for neutron resonance imaging. This chapter is a navigation guide to the rustdoc API documentation, not a standalone physics textbook.
All implementations reference specific sections of the SAMMY manual and SAMMY Fortran source files. See the rustdoc for each module for detailed equations and citations.
Cross-Section Formalisms
| Formalism | ENDF LRF | Module | SAMMY Reference |
|---|---|---|---|
| Reich-Moore | LRF=3 | reich_moore | Manual Sec 2, rml/ |
| Single-Level Breit-Wigner | LRF=1,2 | slbw | Manual Sec 2, mlb/ |
| R-Matrix Limited | LRF=7 | rmatrix_limited | Manual Sec 2 |
| Unresolved Resonance Region | LRU=2 | urr | Hauser-Feshbach |
The penetrability and
channel modules provide the underlying
nuclear physics: hard-sphere phase shifts, penetrability factors, wave numbers,
and statistical spin weights.
Broadening Models
Doppler Broadening
Free Gas Model (FGM) convolution accounting for thermal motion of target nuclei.
- Module:
doppler - SAMMY reference:
dop/module, manual Sec 3.1 - Key function:
doppler_broaden()using psi/chi auxiliary functions on an adaptive grid
Resolution Broadening
Instrument resolution broadening from flight-path uncertainty, timing jitter, and moderator pulse width.
- Module:
resolution - SAMMY reference:
convolution/module, manual Sec 3.2 - Supports: Gaussian convolution, Gaussian + exponential tail, tabulated resolution functions
Transmission Model
Beer-Lambert transmission: T(E) = exp(-sum_i n_i sigma_i(E))
Where n_i is the areal density (atoms/barn) and sigma_i(E) is the broadened total cross-section for isotope i.
- Module:
transmission - SAMMY reference:
cro/,xxx/modules, manual Sec 2, Sec 5 - Handles multi-isotope samples with shared Doppler temperature (one global temperature parameter, optionally fitted jointly with densities)
Fitting Engines
Levenberg-Marquardt
Standard nonlinear least-squares minimization for Gaussian-distributed data.
- Module:
lm - SAMMY reference:
fit/module, manual Sec 4 - Parameters: areal densities with optional bounds, optional temperature fitting
Poisson KL Divergence
Maximum-likelihood fitting for low-count data where Gaussian statistics break down.
- Module:
poisson - Reference: TRINIDI approach (
trinidi/reconstruct.py) - Uses bounds-based preconditioning for joint density + temperature fits
ENDF Nuclear Data
Resonance parameters are sourced from the ENDF/B library via the IAEA API:
- Module:
retrieval-- download and cache - Module:
parser-- parse ENDF-6 File 2 - Module:
resonance-- data structures
Supported libraries: ENDF/B-VIII.0, ENDF/B-VIII.1, JEFF-3.3, JENDL-5.
Further Reading
- SAMMY User's Guide (ORNL/TM-9179/R8)
- ENDF-6 Formats Manual (BNL-203218-2018-INRE)
- ENDF/B-VIII.0 (Nuclear Data Sheets, 2018)
Contributing
Development Setup
git clone https://github.com/ornlneutronimaging/NEREIDS.git
cd NEREIDS
cargo build --workspace
For Python binding development, use pixi:
pixi run build # maturin release build
pixi run test-python # pytest
Pre-Commit Checklist
Run these three commands before every commit:
cargo fmt --all
cargo clippy --workspace --exclude nereids-python --all-targets -- -D warnings
cargo test --workspace --exclude nereids-python
cargo fmtapplies formatting (not just--check)cargo clippytreats all warnings as errorsnereids-pythonis excluded because it requires PyO3/maturin build setup
Branch and PR Workflow
- Create a feature branch from
main - Make changes, commit with GPG signatures (
git commit -S) - Push and open a PR against
main - All PRs go through the review pipeline before merge
The repository uses a single remote (origin = ornlneutronimaging/NEREIDS).
All branches and PRs are pushed directly.
Code Guidelines
Physics Modules
- Implement exact SAMMY physics -- no ad-hoc approximations
- Reference SAMMY source files and equation numbers in doc comments
- Validate against SAMMY's own test cases as ground truth
General
- Validate configuration up-front in public entry points
- Guard NaN with
.is_finite()(NaN bypasses comparison guards) - Guard empty collections explicitly (
.is_empty()) - Use named constants instead of magic numbers
- Prefer
return Err(...)for input validation, notdebug_assert!
Testing
# Rust tests
cargo test --workspace --exclude nereids-python
# Python tests (requires pixi)
pixi run test-python
# Build docs locally
cd docs/guide && mdbook build && mdbook serve
Project Structure
NEREIDS/
crates/
endf-mat/ # Element/MAT lookup tables
nereids-core/ # Core types and constants
nereids-endf/ # ENDF retrieval and parsing
nereids-physics/ # Cross-section physics
nereids-fitting/ # Optimization engines
nereids-io/ # Data I/O (TIFF, NeXus)
nereids-pipeline/ # Orchestration
bindings/python/ # PyO3 Python bindings
apps/gui/ # egui desktop application
docs/
guide/ # mdBook user guide (this site)
adr/ # Architecture decision records
references/ # Reference materials
Useful Commands
| Task | Command |
|---|---|
| Build all | cargo build --workspace |
| Run tests | cargo test --workspace --exclude nereids-python |
| Format | cargo fmt --all |
| Lint | cargo clippy --workspace --exclude nereids-python --all-targets -- -D warnings |
| Build Python | pixi run build |
| Test Python | pixi run test-python |
| Build docs | cd docs/guide && mdbook build |
| Serve docs | cd docs/guide && mdbook serve |
| Build rustdoc | cargo doc --workspace --no-deps --exclude nereids-python |